|
What is PPDB? PPDB is a Plant Proteome DataBase for Arabidopsis
thaliana and maize (Zea mays). Initially PPDB was dedicated
to plant plastids, but has now expanded to the whole plant proteome – hence it
was renamed from Plastid PDB to Plant PDB in November 2007.
Which proteins can be searched? All protein-encoding gene models in the Arabidopsis,
maize and rice, as assembled respectively in http://www.arabidopsis.org/, http://www.maizesequence.org/index.html
and http://rice.plantbiology.msu.edu/, supplemented with plastid and mitochondrial genomes. These are all uploaded in PPDB and are linked to each
other via a BLAST alignment. Thus every predicted protein in both species can be searched for experimental and other
information (even if not experimentally identified).
What information is stored in PPDB? The PPDB stores
experimental data from in-house proteome and mass spectrometry analysis,
curated information about protein function, protein properties and subcellular
localization. Importantly, proteins are particularly curated for possible
(intra) plastid location and their plastid function. Protein accessions
identified in published Arabidopsis (and other Brassicacea)
proteomics papers are cross-referenced to rapidly determine previous
experimental identification by mass spectrometry.
Protein report pages. Each protein has a protein report page
that summarizes all information for that protein stored in PPDB. A few other
databases are hyperlinked for each accession to obtain additional information
for this protein accession. These report pages also provide detailed
information about matched mass spectrometry data and how it maps to the
protein. For individual protein searches, this is the best way to obtain
a comprehensive overview.
How to extract information from PPDB? The PPDB has nine search
functions to extract multiple types of information (output) stored in PPDB for
any accession, by simply choosing a search function and selecting the relevant
‘radio buttons’. Information can all be extracted from the PPDB for individual
accessions or in batch format.
The search functions are:
Search by Accession. (TAIR accessions or ZmGI
accessions).
Search with Protein name. (free text search –
note this can be slow).
Search Proteome Experiments. Select sample
source by species, organ, cell type or subcellular fraction.
Search Comparative Proteomics experiments. Select
particular comparative proteomics experiments.
Search Subcellular Proteomes. Extract proteins
based on their curated location in the cell.
Search Protein Function. Extract all proteins with a
selected function.
Search Proteomics Publications. Find major
proteomics publications and the protein accessions identified.
Search Post-translational Modifications. Find all
proteins with the selected PTM and show the modified peptides.
Search Biochemical Pathways. Find details about
(potential) proteins operating in a selected pathway. Currently only Calvin
cycle.
Extractable protein information (via selection of radiobuttons):
Annotation protein name - from original source (DFCI Gene Index)
or TAIR) and in-house (lab) annotation
Subcellular location - curated (in-house) and
predicted
Biochemical properties and topology - denatured or
native mass, pI, gravy, TMDs, cystein content
Function - Functional PFAM domains and functional
classification (MapMan system)
Mass spectrometry based identification - Mowse score,
# matched peptides and queries, homologues
Sample information - genotype, tissue type, cell
type, organelle, suborganellar fraction
Cross-reference to other studies - proteomics papers,
fluorescence labeling studies
Homologues in other species - Arabidopsis, maize,
rice
Disclaimer:
When using the PPDB, please cite the following reference: Sun Q, Zybailov B, Majeran W, Friso G, Olinares PD, van Wijk KJ. (2008) PPDB, the Plant Proteomics Database at Cornell. Nucleic Acids Res. 2009;37(Database issue):D969-D974. doi:10.1093/nar/gkn654
Contact:
Klaas van Wijk (kv35@cornell.edu) or Qi
Sun (qisun@cornell.edu)
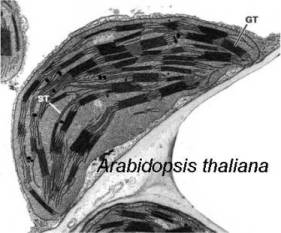
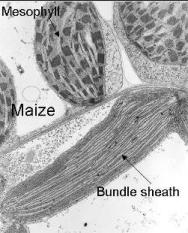
(Image courtesy of Buchanan, B., Gruissem, W. Jones, R.L. Biochemistry
and Molecular Biology of Plants , Publisher - American Society of
Plant Physiologists)
|











